In scientific publishing, the quality of a study is in the details of its findings, methodology, and conclusions. Journal covers offer an artistic snapshot, asking readers to delve deeper. Our work was featured on the covers of Developmental Cell and Cell Host & Microbe.
In our 2021 collaboration with Richard White‘s lab, “Cooperation between melanoma cell states promotes metastasis through heterotypic cluster formation“, we explored how heterogeneous populations of melanoma cells cooperate during metastasis. The cover of Developmental Cell captures this concept. The multicolored dandelion seeds floating in the wind symbolize the collaboration between proliferative (PRO) and invasive (INV) cell states in melanoma, leading to metastasis via heterotypic circulating tumor cell clusters. This collaboration resembles the combined effort of individual dandelion seeds, ensuring the survival and spread of the plant. The ethereal artwork was designed by Wenjing Wu, adding an artistic layer to our scientific findings.
This work was part of Nate Campbell‘s PhD thesis.
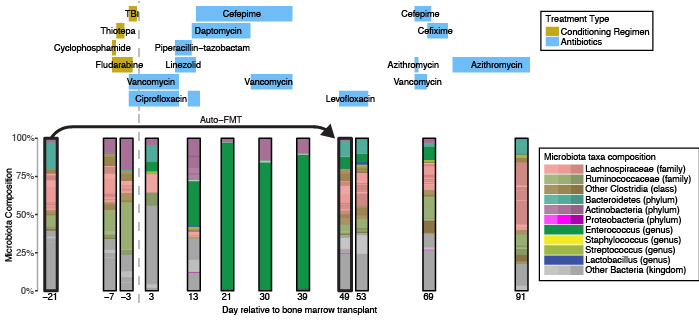
In 2023, our lab made another breakthrough with the paper “The TaxUMAP atlas: Efficient display of large clinical microbiome data reveals ecological competition in protection against bacteremia“. The cover of Cell Host & Microbe shows a topographical representation of our findings. The peak regions in vivid red highlight microbiome states linked to a high risk of bloodstream infection, while the troughs in tranquil blue symbolize states that correlate with a lower risk. This juxtaposition of highs and lows is a helpful visualization that lead us to the ecological competition within the gut microbiome, particularly between the bacterium Klebsiella and other enterobacteria.
This work was led by Jonas Schluter, Ana Djukovic and Brad Taylor.